Cell and Gene Therapy
Single-cell multiomic analysis provides the most precise view of the cellular ecosystem.
Traditional methods for genomic and transcriptomic analysis have been limited by tradeoffs between precision, sensitivity, and resolution. Bulk sequencing lacks the sensitivity and resolution to identify rare variants. Most single-cell sequencing methods either have reduced precision due to error-prone, uneven genome amplification, or provide limited expression level data through RNAseq end counting, which lacks the ability to provide detection of on- and off-target gene edits in non-coding regions or unexpressed genes.
BioSkryb's patented primary template-directed amplification (PTA) directs a high-fidelity polymerase to the genomic DNA template rather than using amplicons as templates. As a result, amplification is pseudolinear, uniform across the genome and between alleles, and subject to minimal error propagation.
Our ResolveDNA® scDNAseq kits and services harness the power of PTA to provide complete, unbiased views of whole genomes or whole exomes from single cells, or other low-input DNA samples.
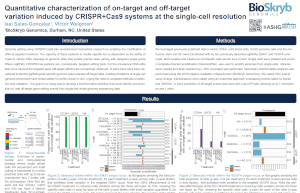
Over 85% of off-target indels detected by ResolveDNA were missed by a predictive algorithm.
For two guide RNAs, ResolveDNA detected hundreds of off-target insertion or deletion mutations (indels) that were not predicted by the Cas-OfFinder algorithm.
Pairing transcriptomic analysis with genomic analysis reveals phenotypic impact.
PTA can also be used for simultaneous single-cell transcriptomic and genomic analysis. Our streamlined ResolveOME™ workflow allows the transcriptome and genome of single cells to be sequenced in parallel, revealing critical information about the potential phenotypic impact of genomic variants.
For cell and gene therapy research and development, the unique capabilities of ResolveDNA and ResolveOME enable identification of rare but potentially significant cellular subpopulations via sensitive, precise detection and cellular attribution of:
- On- and off-target gene edits including SNVs, indels, and larger structural aberrations
- Therapeutic transgene integration events
- Transcript splicing variants
- Gene expression changes associated with altered cell state and/or oncogenic potential
Benefits of ResolveDNA and ResolveOME
- >95% genomic coverage
- Superior coverage uniformity
- Unparalleled sensitivity and precision in calling both SNVs and CNVs
- >95% transcript length coverage
- Ability to link gene expression changes to genomic variation
Confidently Evaluate the Safety and Efficacy of Cell and Gene Therapies
Current approaches to characterizing cell and gene therapies may miss off-target genomic changes and correlated gene expression changes. Read our perspective on how emerging single-cell multiomics technologies put more comprehensive safety and efficacy data in researchers' hands.
Additional Resources
“BioSkryb’s superior PTA technology and exceptional service have been invaluable to our efforts to characterize on- and off-target editing activity of CRISPR-based gene editing technologies. Their team is responsive, swift, and consistently goes above and beyond, delivering high-quality single-cell whole-genome amplification (WGA) results that outperform competitors across key metrics. The high-quality WGA results delivered by the BioSkryb service team have made it possible to measure single-nucleotide variants (SNVs) at single-cell resolution with higher confidence than ever before.”
Jeffrey Quinn
Assoc. Director, Experimental Genomics & Off-Target Biology, Beam Therapeutics